"Basepaws continues to expand its one-of-a-kind genomic and oral microbiome database with the help of our dedicated research community. Together, we are pioneering new discoveries that will improve the health and well-being of pets everywhere."

Basepaws collaborates with an amazing community of pet parents, veterinary professionals, and industry partners on groundbreaking research that supports the development of affordable, easy to use screening tools for the early recognition and treatment of disease.

We collect clinical data through partnerships with veterinary professionals and university researchers. This allows us to obtain samples from dogs and cats that match particular study criteria.

We have a dedicated community of pet parents who fill out our health history survey and agree to have their pets’ samples included in our citizen science studies.
Our science team conducts follow-up work with our active and loyal community of pet parents to match pet samples in our database with veterinary medical records that confirm diagnoses and health histories.
"Basepaws continues to expand its one-of-a-kind genomic and oral microbiome database with the help of our dedicated research community. Together, we are pioneering new discoveries that will improve the health and well-being of pets everywhere."

Basepaws research aims to understand the genetic and oral microbiome factors associated with a pet’s increased risk for disease. Our studies directly inform the development of at-home screening tools that detect signs of disease sooner and help pets get the individualized care they need. We focus on the following seven areas: Oral Health, Renal and Urinary Diseases, Gastrointestinal Diseases, Endocrine Diseases, Dermatological Diseases, Cancer, Longevity.
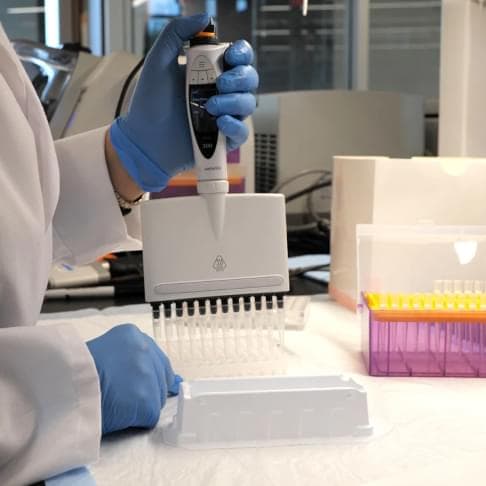


Low-Pass Whole Genome Sequencing (WGS) with Computational Imputation

Next Generation Amplicon Sequencing (NGS)

Oral Microbiome Analysis